Partitioning US counties with migration data
Published:
The IRS provides yearly migration flows between counties in the United States based on where people file their taxes in consecutive years.
Using this data, we want to know whether there are sets of counties in the US that do not have migrants traveling between them. More specifically, we want to find three sets of counties: \(A\), \(B\), and \(C\), such that a.) there are no migrations over a certain period of time between any counties in \(A\) or \(B\) (i.e. there are no migrants that travel between counties \(u\) and \(v\) for all \(u \in A\) and all \(v \in B\)); b.) the size of \(C\) is as small as possible; and c.) the sizes of \(A\) and \(B\) are balanced (for some definition of balanced). This is the vertex separator problem and is known to be NP-Hard [1,2].
In this notebook we formulate this problem as an mixed integer program (MIP), solve an instance of the program for migrations over 5 years in the US, and then map the results to get some insight about the structure of migration in the US. To do this we use the migration-lib package to interface with the IRS migration data, the pulp package to create the MIPs for CPLEX (a commercial solver) to solve, and the simple-maps package to map our results.
[1]. Feige, Uriel, Mohammad Taghi Hajiaghayi, and James R. Lee. “Improved approximation algorithms for minimum weight vertex separators.” SIAM Journal on Computing 38.2 (2008): 629-657. link
[2]. Kim, Mijung, and K. Selçuk Candan. “SBV-Cut: Vertex-cut based graph partitioning using structural balance vertices.” Data & Knowledge Engineering 72 (2012): 285-303. link
Setup packages
%matplotlib inline
import sys,os, time
import numpy as np
import pulp
import re
sys.path.append(os.path.join(os.getcwd(),"migration-lib"))
sys.path.append(os.path.join(os.getcwd(),"simple-maps"))
import MigrationDataUSA
from simplemaps.SimpleFigures import simpleMap,simpleBinnedMap
Load migration matrices
Our migration data is in the form of a list of \(m\) migration matrices, \(\{T_1, T_2, \cdots, T_m\}\), where \(m\) is the number of years for which we have data. Each migration matrix \(T^{(t)}\) is of size \(n \times n\), where \(n\) is the number of counties in the US. A particular entry in one of the migration matrices, \(T^{(t)}_{ij}\), represents the number of people that travel from county \(i\) to county \(j\) in the \(t^{th}\) year of data.
The first step of our method is aggregating the migration matrices for all the years that we care about: \(T = \sum_{t=1}^{m} T^{(t)}\). Now an entry will represent the total number of migrants that travel between two locations over all the years of data for which we care about.
startYear, endYear = 2009, 2014
years = range(startYear, endYear + 1)
numYears = len(years)
migrationMatrices = MigrationDataUSA.getMigrationMatrixRange(startYear,endYear+1)
for matrix in migrationMatrices:
np.fill_diagonal(matrix, 0.0)
countyList = MigrationDataUSA.getCountyList()
numCounties = len(countyList)
aggregateMigrationMatrix = np.sum(migrationMatrices, axis=0)
print aggregateMigrationMatrix.shape
(3106, 3106)
shapefileFn = "simple-maps/examples/cb_2015_us_county_500k_clipped/cb_2015_us_county_500k_clipped.shp"
shapefileKey = "GEOID"
Method to generate MIP for solving the Vertex Separator problem
For our implementation of the Vertex Separator problem we need to have some definition of what “balanced” sets are. We will introduce a parameter \(\alpha\) and constrain the sizes of \(A\), and \(B\) to be greater than \(\alpha n\). Valid values of \(\alpha\) will be in the range \([0, 0.5)\).
Our definition of the problem is as follows:
\[\displaylines{ \max_{A,B} \sum_{i=1}^{n} A_i + \sum_{i=1}^{n} B_i \\\ \text{s.t.} \\\ \\\ A_i + B_i \leq 1 \;\;\;\;\;\; \forall i \in \{1, \cdots, n\} \;\;\;\text{(Constraint 1)}\\\ A_i + B_j \leq 1 \;\;\;\;\;\; \forall i,j \in \{1, \cdots, n\} \text{ where } T_{ij}>0 \;\;\;\text{(Constraint 2)}\\\ A_j + B_i \leq 1 \;\;\;\;\;\; \forall i,j \in \{1, \cdots, n\} \text{ where } T_{ij}>0 \;\;\;\text{(Constraint 3)}\\\ \sum_{i=1}^{n} A_i \geq \alpha n \;\;\;\text{(Constraint 4)}\\\ \sum_{i=1}^{n} B_i \geq \alpha n \;\;\;\text{(Constraint 5)}\\\ A_i \in \{0,1\} \;\;\;\;\;\; \forall i \in \{1, \cdots, n\} \;\;\;\text{(Constraint 6)}\\\ B_i \in \{0,1\} \;\;\;\;\;\; \forall i \in \{1, \cdots, n\} \;\;\;\text{(Constraint 7)} }\]Here \(A\) and \(B\) are lists of size \(n\) of binary decisions variables. An entry \(A_i = 1\) means that county \(i\) belongs to set \(A\), similarly an entry \(B_i = 1\) means that it belongs to set \(B\). We define \(C\), the cut set, as all counties \(i\), for which \(A_i = 0\) and \(B_i=0\). Constraint 1 will ensure that a county can not belong to both sets \(A\) and \(B\). Constraints 2 and 3 ensure that atleast one county, from each pair of counties that has migrants travelling between them, will belong to set \(C\). Constraints 4 and 5 will ensure the balance defintion that we describe above. Finally, constraints 6 and 7 are the binary constraints on \(A\) and \(B\).
def vertex_separator(T,alpha=0.25,timeLimit=60):
assert len(T.shape) == 2, "Input must be a matrix"
assert T.shape[0] == T.shape[1], "Input matrix must be square"
n = T.shape[0]
vertices = range(n)
prob = pulp.LpProblem("VertexSeparator", pulp.LpMaximize)
u = pulp.LpVariable.dicts('u', vertices, cat = "Binary")
v = pulp.LpVariable.dicts('v', vertices, cat = "Binary")
prob += pulp.lpSum([u[i]*1.0 for i in vertices] + [v[i]*1.0 for i in vertices]), "Objective"
#Least absolute deviations
for i in vertices:
prob += u[i] + v[i] <= 1, "Node %d can not be in both clusters" % (i)
for i in vertices:
for j in vertices:
if T[i,j] > 0:
prob += u[i]+v[j] <= 1, "Edge %d %d constraint 1" % (i,j)
prob += v[i]+u[j] <= 1, "Edge %d %d constraint 2" % (i,j)
prob += pulp.lpSum([u[i] for i in vertices]) >= np.floor(alpha * n), "First cluster size constraint"
prob += pulp.lpSum([v[i] for i in vertices]) >= np.floor(alpha * n), "Second cluster size constraint"
cplexOptions = [
"set timelimit %d" % (timeLimit)
]
solver = pulp.solvers.CPLEX_CMD(
path="/opt/ibm/ILOG/CPLEX_Studio1261/cplex/bin/x86-64_linux/cplex",
msg=1,
keepFiles=1,
mip=True,
options=cplexOptions
)
print "Starting solver with time limit of %d seconds" % (timeLimit)
prob.solve(solver=solver)
print "Status:", pulp.LpStatus[prob.status]
new_u = np.zeros((n,))
new_v = np.zeros((n,))
if pulp.LpStatus[prob.status] == "Optimal":
print "Objective function: %f " % (pulp.value(prob.objective))
for v in prob.variables():
varName = v.getName()
value = pulp.value(v)
if varName.startswith("u_"):
vertexIndex = int(varName.split("_")[1])
new_u[vertexIndex] = value
elif varName.startswith("v_"):
vertexIndex = int(varName.split("_")[1])
new_v[vertexIndex] = value
else:
print varName
else:
raise ValueError("Solve failed")
return new_u, new_v
Solve an instance of the problem
Here we solve an instance of the problem for \(\alpha=0.2\). This means that the solution is constrained such that the size of both \(A\) and \(B\) have to each be greater than \(20\%\) of the total number of vertices.
startTime = float(time.time())
alpha = 0.3
try:
A, B = vertex_separator(aggregateMigrationMatrix, alpha=alpha, timeLimit=540)
print "Size of cluster 1: %d" % (np.sum(A))
print "Size of cluster 2: %d" % (np.sum(B))
C = (A==0) & (B==0)
print "Size of vertex cut: %d " % (np.sum(C))
assert np.sum(A) + np.sum(B) + np.sum(C) == numCounties
except ValueError as e:
print e
pass
print "Finished in %0.4f seconds" % (time.time()-startTime)
Starting solver with time limit of 540 seconds
Status: Optimal
Objective function: 2881.000000
Size of cluster 1: 931
Size of cluster 2: 1950
Size of vertex cut: 225
Finished in 567.5502 seconds
Display results
viz = np.zeros_like(A)
for i in range(len(countyList)):
if A[i] == 1:
viz[i] = 1
elif B[i] == 1:
viz[i] = 2
else:
viz[i] = 0
sizeA = np.sum(A)
sizeB = np.sum(B)
sizeC = np.sum(C)
labels = [
"Cut ($|C|=%d$)" % (sizeC),
"Cluster 1 ($|A|=%d$)" % (sizeA),
"Cluster 2 ($|B|=%d$)" % (sizeB)
]
simpleBinnedMap(
shapefileFn,
shapefileKey,
dict(zip(countyList, viz)),
labels = labels,
title="Solution for $\\alpha=%0.2f$" % (alpha),
)
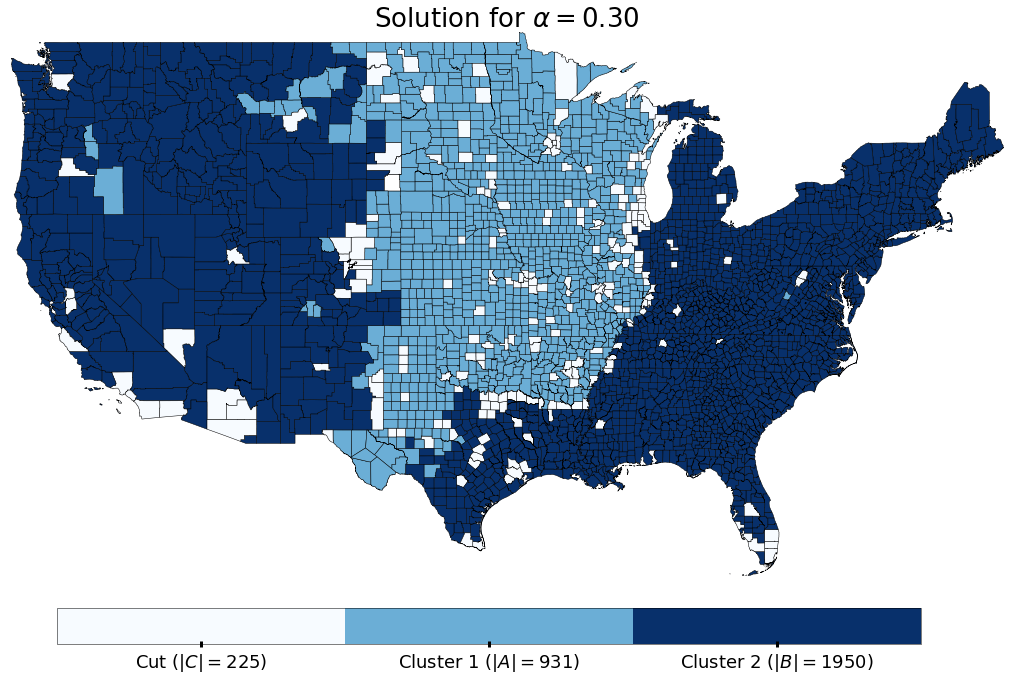
From these results we can see that there is a group of 931 counties (colored in light blue), largely in the mid western part of the United States, and another group of 1950 counties (colored in dark blue), that do not exchange migrants, at all, over the 5 years between 2009 and 2014. The individuals in these sets of counties do migrate to/from the smaller “cut set” of counties (the 225 counties colored in white), however there is not a single observed migration between the larger two sets. In the terms of the Vertex Separator problem, if we were to remove the counties colored white, then the two remaining portions of the country would be cut off from each other.
